How DNA could store all the world’s data in a semi-trailer

Roula Khalaf, Editor of the FT, selects her favourite stories in this weekly newsletter.
Last October, Deep Purple’s “Smoke on the Water” and Miles Davis’ “Tutu” made history again — not musical this time but scientific. Researchers from the University of Washington, Microsoft and Twist Bioscience successfully stored these two recordings on to a DNA sequence that could be decoded and played back without any loss of quality.
It was the latest sign that storing data on DNA is becoming a serious prospect. Scientists have long seen DNA as a potential storage medium because it can store large amounts of data in a small space and is stable over thousands of years. In 2012, scientists at Harvard managed to store the text of a book on DNA, and in early 2017 the Defense Advanced Research Projects Agency’s (Darpa), the US government branch dedicated to national security research, announced plans to fund further research into DNA storage.
If successful, it could be the answer the problems of archiving the increasing amounts of information the world is creating. Data storage is a bigger problem than many people appreciate. Most digital archives — from music to research — are currently saved on magnetic tape, explains Gurtej Sandhu, senior fellow at Micron Technology, a digital memory products company with a near-$50bn market cap.
Like a cassette tape, only “much bigger and more high-tech”, magnetic tape is the cheapest way to store data. With content increasing exponentially, however, “tapes are going to fill up hundreds of warehouses,” Mr Sandhu says. “And the tape doesn’t store forever, you have to rewrite it — the information will disappear after 10 years.”
DNA, however, is very good for storing and copying information, as our own bodies prove. “[Whether you are] a virus, a cucumber, an elephant, Donald Trump, whatever,” says Yaniv Erlich, a computer scientist at Columbia University who published a paper on DNA storage in March. “You store the most important information in your life in your DNA.”
The genetic molecule is extremely small, so encoding data into it would solve the tape real-estate problem. “We need about 10 tons of DNA to store all the world’s data,” adds Mr Erlich. “That’s something you could fit on a semi-trailer.”
DNA also has longevity. “DNA has been around for the last 3.5bn years,” points out Mr Erlich. “It’s not going to be obsolete even in 10,000 years.”
Emily Leproust, chief executive at Twist Bioscience, says DNA is cheap and quick to copy. “It takes $1 and one hour to copy a tube of DNA,” says Ms Leproust. “It may sound high — but if you have in a tube the equivalent of a data centre, you can copy an entire data centre for $1 and one hour. That is absolutely unheard of.”
Translating the binary code which makes up digital data into the chemical rungs that form DNA spirals is less complex than might be expected, according to Sriram Kosuri, assistant professor of chemistry and biochemistry at the University of California, Los Angeles.
Prof Kosuri co-authored a 2012 paper — which he says was “probably one of the simplest science papers ever published in some ways” — showing that the 0s and 1s of binary code can be translated into pairings of DNA bases, A, T, C, G. So A could represent 00, T might represent 01, and so on.
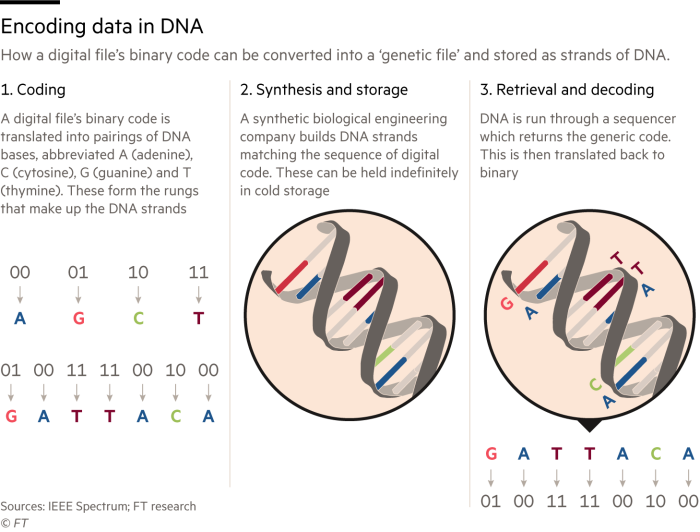
A DNA strand can then be built matching the sequence of digital code. To read the data, the DNA is run through a sequencer and decoded.
Synthesising DNA, however, can be messy — not all the molecules make it and sometimes bits repeat. To reduce errors when reading back, Mr Erlich and his team came up with an algorithmic solution akin to Sudoku maths puzzles. In Sudoku, “even if you lose some of the hints you can still solve the puzzle,” says Mr Erlich.
Using the Sudoku technique, Mr Erlich and his team were able to store information at a density of 215 petabytes in one gram of DNA, and read it back accurately. One petabyte is equivalent to 900bn pages of plain text. Darpa has funded Mr Erlich to develop the system further.
Computing architects and synthetic biologists are now designing systems to automate the DNA storage processes, and let you search the DNA for specific files.
Luis Ceze, a computer sciences professor at the University of Washington, explains that the process involves wet parts — the suspended DNA — and electronics. “All of the liquid, fluidics part has to be automated,” he said. “You’re not going to employ 100m people in data centres to move DNA around.”
Cost remains an obstacle. While reading DNA sequences is now relatively cheap and easy, writing DNA can be prohibitively expensive. Twist Bioscience charges between 7 and 9 cents per DNA base; storing 12mb of data on DNA costs around $100,000.
Robert Carlson, a consultant at the biological engineering company Biodesic, says that for a DNA drive to compete with a single tape drive, it would need to read and write “the equivalent of around 10,000 human genomes a day”. At present, he estimates that the global DNA synthesis industry writes just three human genomes-worth of DNA per day.
Ms Leproust, whose company uses 3D printing techniques to write DNA, is optimistic, however. “We believe that we have a road map to be able lower the cost of DNA (synthesis) by a millionfold,” she says, although she concedes that the timeframe for achieving this goal is still unclear.
50 ideas to change the world
We asked readers, researchers and FT journalists to submit ideas with the potential to change the world. A panel of judges selected the 50 ideas worth looking at in more detail. This third tranche of 30 ideas (listed below) is about new ways to handle information and education. The next 10 ideas, looking at advances in healthcare, will be published on March 5, 2018.
- Holograms
- Scanners to read our minds
- Quantum computing
- All the world’s data stored on DNA
- The next challenges for AI
- Rethinking our sleep schedule
- Personalised learning
- Neural networks allow us to read faces
- Learning to overcome digital distractions
- Robots in the classroom
This story has been updated to clarify that Mr Erlich and his team stored information at a density of 215 petabytes in one gram of DNA. The actual amount of data stored was a few magabytes.

Comments